The project
ESCOGITARE is the project in which noze collaborated with ISTI — the CNR Institute of Information Science and Technologies “A. Faedo” in Pisa — to build a system for the creation and sharing of bioinformatics experiments, aimed at a community of biologists distributed across Italy.
The goal of the system is to give researchers and labs a web tool where they can:
- compose scientific workflows visually (without writing code)
- run them on a pre-existing computational grid
- share both the workflows and the results with the rest of the scientific community
- publish information about projects, experiments, data banks, news and related materials
Results were presented at the closing conference on 5 February 2010 in Pisa at ISTI-CNR.
The consortium
- ISTI-CNR Pisa — Institute of Information Science and Technologies “A. Faedo”
- CRA — Consiglio per la Ricerca e la sperimentazione in Agricoltura, with its two institutes involved in the project:
- ISSDS-CRA Florence — Experimental Institute for Soil Study and Conservation
- ISC-CRA Fiorenzuola — Experimental Institute for Cereal Research
- Research Center on Software Technology
- noze — portal, visual interface and grid integration
Following the 2015 reform, CRA and its institutes were merged into CREA — Council for Agricultural Research and Agricultural Economics Analysis, which today carries forward the research mission of the project’s agricultural partners.
noze’s role
noze took care of three main things, across the entire stack:
1. Interfacing the existing grid
ISTI-CNR already had a working computational grid (based on Globus Toolkit, Active BPEL and MDS). noze interfaced it with a scientific workflow management system, so that experiment administration, deploy and run operations became accessible from the web portal — without requiring grid sysadmin skills from the end users. Among the administration functions exposed: shutting down Globus containers, deploy/undeploy GAR, querying the service index, removing machines, stop IP-forwarding.
2. Visual Workflow Designer
The most distinctive component built by noze: a drag-and-drop visual editor to compose scientific workflows that can be shared between researchers. The dynamic palette contains specialised bioinformatics macroblocks (FBClustalW, FBCondition, FBHuman, FBMPIBlast, FBPrimer3, FBReceiveSequence, FBSequence2Primer3Adapter), reusable and configurable. The workflows drawn visually are translated automatically to BPEL and executed by the Active BPEL Engine on the Globus grid.
Design goal: keep the editor as simple as possible, so it could be used by a typically non-IT audience (biologists and lab researchers), without having to write code or BPEL by hand.
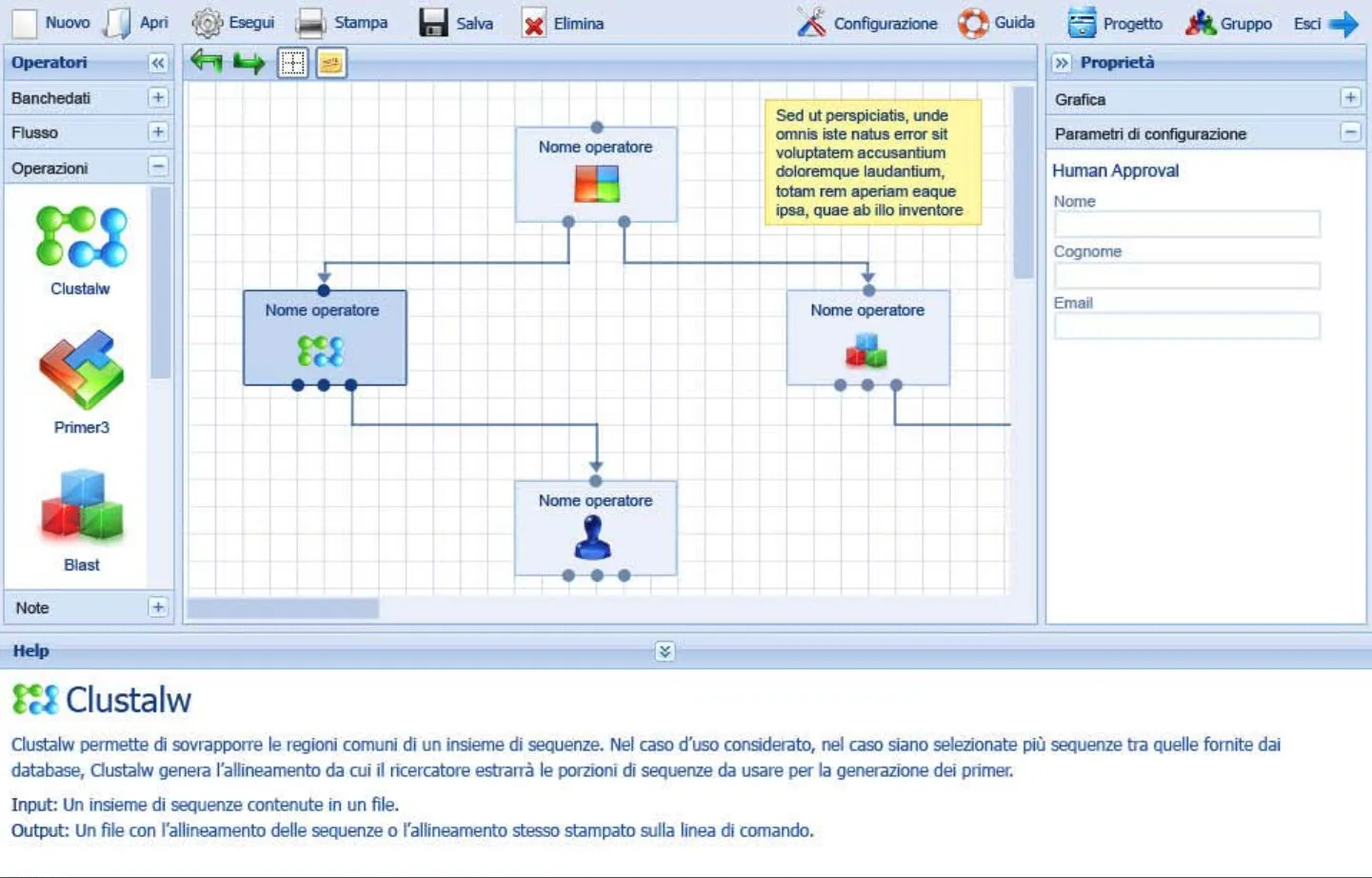
Screenshot of the Visual Workflow Designer: on the left the operators palette (ClustalW, Primer3, Blast), in the center the canvas with the connected blocks composing a bioinformatics workflow, on the right the configuration-parameters panel, at the bottom the contextual help.
3. Web 2.0 CMS and portal
noze integrated the entire workflow management system with a CMS in charge of publishing the content related to bioinformatics projects and experiments (Projects, Research, Data Banks, Forum, AccessGrid, News & Events sections).
The CMS visual style was designed to integrate with the rest of the portal — the Visual Workflow Designer, the grid administration interface and the project/workflow management interface — so that the user perceives a single coherent environment, not the aggregation of separate tools.
Web 2.0 approach and usability
The portal’s visual design follows the Web 2.0 principles — an approach to the internet that combines visual, functional and collaborative aspects — because the goal of the system is precisely to enable a large community of biologists to compose and share scientific workflows.
In addition to the computation grid, the portal also supports AccessGrid for multi-site scientific videoconferencing, completing the distributed-collaboration experience.
Bioinformatics use case
A typical “in silico” experiment implemented on the platform:
- Search for sequences via a connection to GeneBank
- Multi-sequence alignment via ClustalW
- Design of the most appropriate primers via Primer3
- Specificity verification of sequences via BLAST
- Lab tests and verification of the results
The whole workflow is composed visually in noze’s editor, executed on the Globus grid via BPEL, and shared with the community through the portal.
